I.
Writing is an act of discovery. Sometimes you sit down, intent on making a specific argument, only to find (upon deeper research) that the argument is garbage and doesn’t hold up to scrutiny! Writing has a beautiful tendency to reveal holes in one’s logic, as it did for this essay.
Initially, the goal was to make a long argument about how polymerase chain reaction, or PCR, hasn’t changed basically at all since 1987, when the first “modern” thermocycler machines were released. The thesis was that there must be many ways to make PCR significantly faster, cheaper, better. This argument is roughly correct, but not to the degree anticipated. Time-savings with new PCR technologies are modest, and scientists are reluctant to buy cheap PCR machines for a few reasons (more on that later).
And yet, this essay still seemed worth writing. The inspiration stemmed from two ideas submitted for the Fast Biology Bounties — from Sebastian Cocioba and “Utah” Hans — about plans to create photonic PCR machines. The gist of photonic PCR is that you can use LED lights or lasers to rapidly heat samples, thus running 40 cycles of DNA amplification in the span of 6 minutes. I gave out $3,500 in microgrants to support these ideas, courtesy of Astera Institute.
Faster PCR may not seem like a particularly desirable problem to work on. But it’s important for a few reasons. First, even a marginal improvement to a widespread method can have huge downstream effects on scientific productivity as a whole. And second, as more biology experiments get automated, and robots run these experiments 24/7, those time improvements will scale exponentially. Trimming 20 minutes off PCR may not be a big deal to human scientists, but it might matter a lot to robots!
It turns out, though, that many of my (weakly held) assumptions were quickly overturned. As exciting as photonic PCR seems, I think it’s unlikely to show up in academic labs anytime soon.
II.
PCR is a decades-old method to copy DNA. Millions of biologists use it nearly every week for everything from cloning genes to diagnosing diseases. The way it works is fairly simple, too. Just take a little tube and add a DNA sequence, or the molecule to be copied. Next, add a DNA polymerase enzyme and some primers, which are short DNA snippets that bind to the DNA sequence to be copied. And finally, sprinkle in some nucleotides (the raw building materials for DNA) and magnesium. Finally, place the tube inside a machine, roughly the size of a DVD player, called a thermocycler, the sole job of which is to ramp temperatures up and down, again and again.
The machine starts by increasing the temperature to about 96°C, which melts the double-stranded DNA molecule, breaking it into two separate pieces. Then, the temperature drops to 60°C, the perfect temperature for primers to latch onto those DNA molecules. Next, the device goes back up to about 70°C, the temperature at which the polymerase enzyme is most active. Each polymerase seeks out a primer, grabs onto it, and begins copying the DNA by stitching together nucleotides. And finally, the thermocycler goes back to 96°C to begin the cycle anew. Every time this cycle runs, the amount of DNA in the tube roughly doubles. There are typically 30 cycles in a single PCR experiment, which means that two strands of DNA become (assuming perfect efficiency) 230 copies, or 1.07 billion molecules.
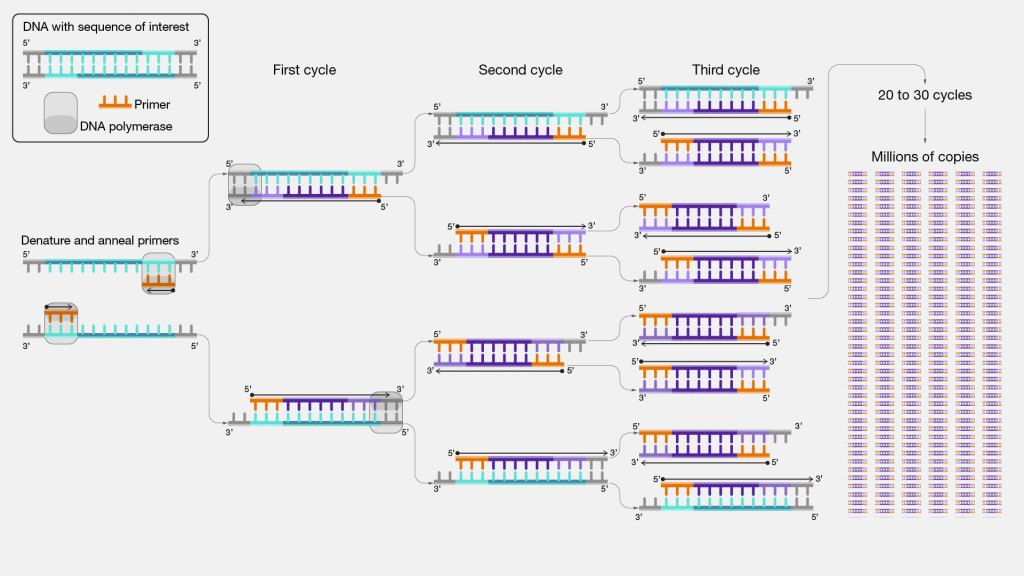
How PCR works. From genome.gov.
Running a PCR today takes about an hour. Each cycle takes a few minutes. Why can’t this happen much faster?
There are three factors involved: diffusion, or the time it takes for molecules to move to their correct locations, the length of the DNA being copied, and the ramp rates between each temperature. Diffusion matters because DNA cannot be copied until the primers and polymerase have “found” their targets; longer DNA strands take more time to copy because polymerase moves at a specific pace (about 60 nucleotides per second); and ramp rates just mean thermocyclers must cool from, say, 96°C to 70°C. All of this takes time.
With these factors in mind, though, we can start thinking about ways to cut down on these bottlenecks.
The easiest way to speed up PCR is to reduce the number of cycles. Most scientists use 30 cycles as a default, but is there a particular reason for this? Presumably, some ingredients in the PCR mix (like primers, or nucleotides) would run out before cycle 30, and thus the final few cycles wouldn’t actually mean much anyway.
But not so! While some ingredients in the PCR mix get reused between cycles (like the polymerases), others are consumed every time DNA is made. Primers are physically incorporated into new strands, for example, and nucleotides are used to build them. But none of these ingredients run out before cycle 30. A 50-microliter PCR tube has about 6 × 10¹⁵ molecules of each nucleotide, enough to make over 10¹² copies of a 3,000-base gene. (For context, 10¹² is 1,000-times larger than 230) Primers and polymerases aren’t limiting either. Polymerases only start to become scarce after about 35 cycles.
The actual limitation in PCR is “product reannealing,” which just means that as the DNA copies become more abundant (in the hundreds of millions of copies) they start to find each other during the annealing step faster than the primers can bind them. As the amplified DNA starts to compete with primers, each cycle gets less efficient. Cutting a PCR from 30 to 25 cycles would just reduce the DNA yields, then, which would also decrease the likelihood that experiments work well downstream. Therefore, I’m not convinced that reducing the number of PCR cycles is a good way to save time.
The second way to speed up PCR is to make faster polymerases. Taq, a polymerase isolated from Thermus aquaticus, a “heat-loving” bacterium found in near-boiling water in Yellowstone National Park, copies DNA at about 60 nucleotides per second. It could theoretically replicate a 3,000-base DNA sequence in less than a minute. In reality, though, the polymerase “falls off” the DNA all the time (like, every second) meaning it actually goes much slower. (New England Biolabs, the company that sells Taq, recommends one minute of extension time per kilobase of DNA, or 3 minutes for 3,000 bases.) The DNA extension step at 70°C is by far the longest part of PCR.
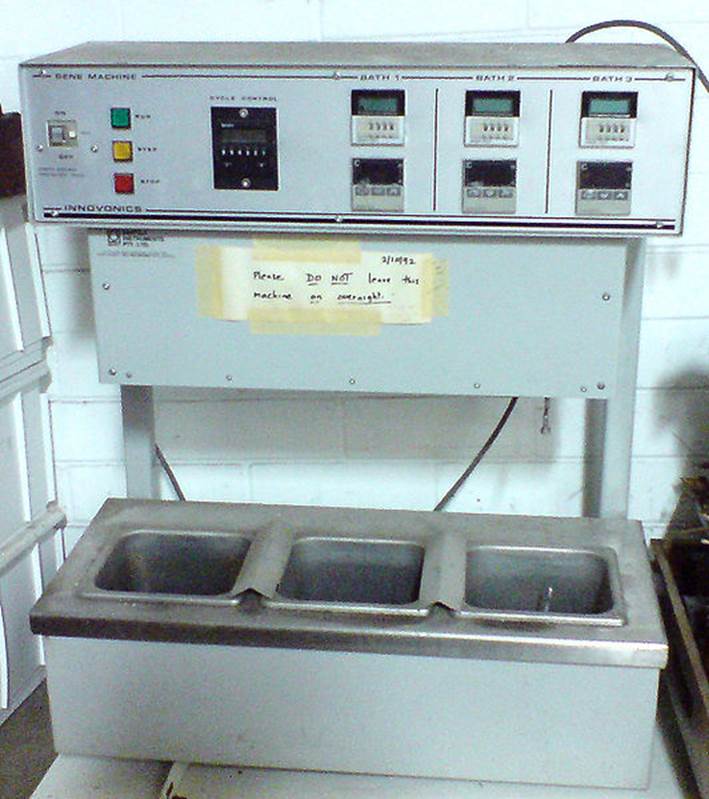

Before Taq was discovered, scientists were forced to add “fresh” polymerase to their tubes during each cycle, manually moving them between water baths held at the three different temperatures.
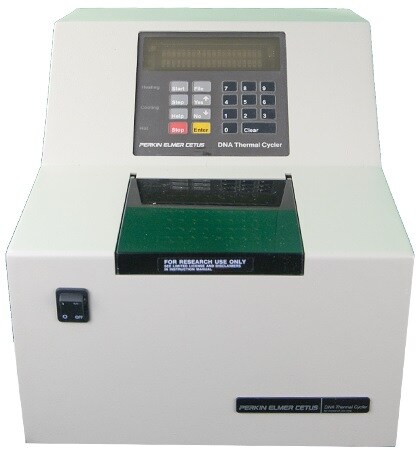

The first “modern” thermocycler, the TC1, was released in 1987 by Perkin-Elmer and Cetus.
Other polymerases, like Phusion, are much faster; not because they move faster along the DNA, but because they don’t fall off so much. Phusion is an engineered protein made by attaching a Pyrococcus polymerase to a small DNA-binding protein, Sso7d, that makes it hold onto DNA strands more tightly. ThermoFisher, the company behind Phusion, recommends just 15 seconds per kilobase for extention. In other words, swapping out one polymerase for another can save more than an hour of time on PCR. This is already an incredible time savings, and I’m skeptical we’ll find anything besides marginal time savings by engineering faster polymerases.
Finally, we can speed up PCR by making faster thermocyclers. Modern machines are slow and outdated. A few years ago, ThermoFisher bought a bunch of thermocyclers and directly compared how fast they heat and cool samples. Across the instruments, the average upward ramp rate was 4.75 °C/s and the average cooling rate was 3.82 °C/s. Cheaper machines, like the Bio-Rad T100, have a ramp rate closer to 2.5°C/s (and these are the most common devices in research labs.) In practice, then, ramping temperatures eats up a lot of time! Every time the machine drops from 95°C to 60°C, it takes about 10 seconds.
Thermocyclers have not changed much in the last forty years, either. They usually have a big metal block — aluminum, or silver if you pay extra — with holes drilled into it to hold the tubes. A lid comes down and sandwiches heating elements on top of the block, and a Peltier element heats the block by running electrical current to warm one side while cooling the other. To switch temperatures, the machine reverses the electrical current and fans blow the extra heat away. The ramp rates are slow because the machine is fighting the thermal mass of a big chunk of aluminum; it resists sudden temperature changes.
The only real option left to speed up PCR, then, is to massively speed up ramp rates. Cutting cycles doesn’t really work, and polymerases have already been so beautifully optimized that further gains would be tiny. But lasers and LED lights could, perhaps, make ramp rates nearly instant. How might that work?
III.
People have been using LEDs and lasers to do PCR for decades, though I’m not aware of any designs that have been commercialized. The gist is that, rather than heating a big aluminum block, you can shine a light directly at a droplet of liquid to heat it up. If the droplet is sufficiently small, with a high surface area-to-volume ratio, that heat will quickly dissipate and cool the droplet once the light is switched off. (No fans required.)
For a 2009 paper, researchers made tiny droplets of PCR mix and dropped them into mineral oil to prevent evaporation. They placed the droplets onto a glass slide held at 58°C, the temperature used for both annealing and extension. Then, they aimed an infrared laser overhead and heated the droplets to 93°C for the denaturation step. Water absorbs infrared light strongly, whereas mineral oil absorbs about 40x less, so the laser heats the water and nothing else. As soon as the laser switches off, heat quickly drains from the water into the surrounding oil.
In one experiment, they used this setup to amplify a 187-basepair strand of DNA and finished 40 cycles in about six minutes. Because the volumes were so small, and the ramp rates so fast, each cycle only needed two seconds for denaturation, one second for primer annealing, and seven seconds for extension! (The extension step alone took about two-thirds of the total time.) The power needed to run the laser is also tiny; it consumes about 3,000-times less energy than a light bulb.
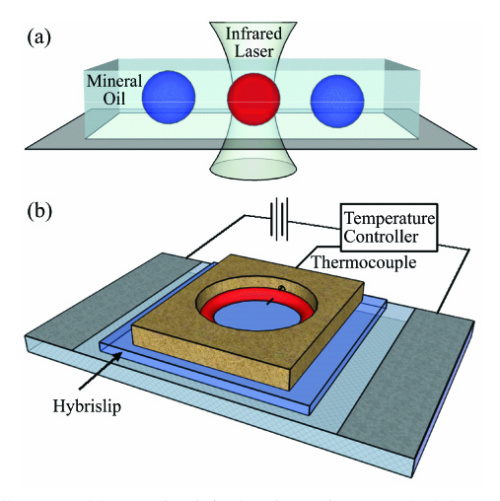
Using an infrared laser to heat up small droplets for PCR. From the 2009 paper.
In 2015, a UC Berkeley team published a similar idea, but used LEDs and a thin gold film instead of a laser. They coated a small piece of plastic with the gold film and dropped the liquid on top. Then, they placed blue LEDs underneath, using them to heat the gold and, in turn, the liquid above it. (Gold is ideal because it absorbs light efficiently and is also inert, so it doesn’t mess up the PCR.) With this simple setup, they could run 30 cycles of heating and cooling, switching between 55 and 95°C, in about six minutes. The ramp rate was about twice as fast as any commercially-available thermocycler.
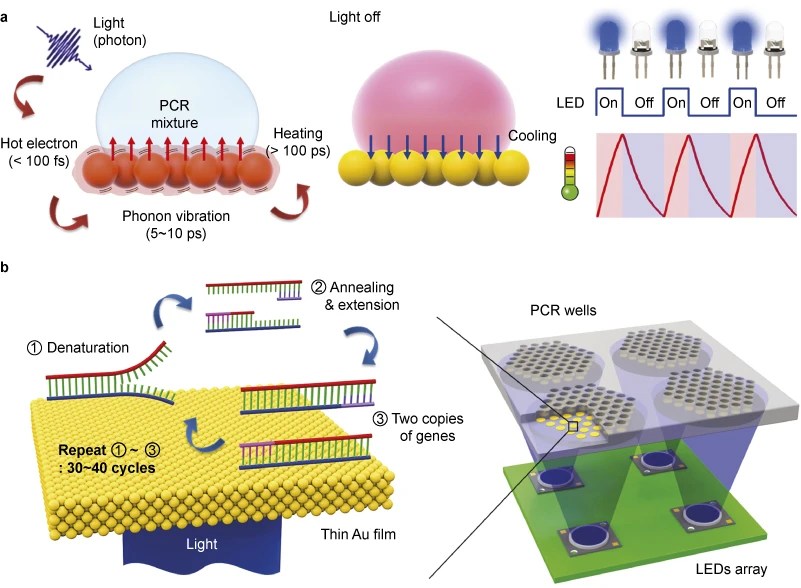
Using gold film and LEDs to do PCR. From the 2015 paper.
The two microgrants that I gave out were for ideas based directly on these prior studies. Sebastian Cocioba is building a small device that uses black PCR tubes and a small laser to heat samples, while “Utah” Hans is (mostly) duplicating the 2015 study, using gold and LED lights to build a PCR machine for less than $50.
Although I was initially excited by these ideas, I’m no longer convinced that photonic PCR will be widely adopted (even if cheap prototypes became available.) My rationale comes from a simple calculation. Imagine using the Phusion enzyme to amplify a gene made from 3,000 bases of DNA, running 30 cycles of amplification at the temperatures mentioned before: 98°C (denaturation, 10 seconds) → 60°C (annealing, 20 seconds) → 72°C (extension, 45 seconds). How long would this take with “average” ramp rates from commercially-available machines? And how long would it take if the ramp rates were instantaneous instead?
There’s not a big difference between the two! In this scenario, the temperature ramping alone takes about 20 seconds per cycle, or 10 minutes total, and eats up about 20 percent of the total PCR time. So een if the ramping were instant, it wouldn’t matter much; PCR would go from a one-hour thing to a 50-minute thing. This was surprising to me, and I think it partly explains why photonic PCR devices haven’t caught on.
IV.
So photonic PCR won’t save much time. But suppose someone actually built a working photonic PCR machine and sold it for $150. (Utah has sent me a bunch of videos and has a working prototype that cost just $48 to build.) Would scientists buy it?
On paper, yes! A normal thermocycler costs anywhere from $2,500 (on the low end) to $10,000 (with lots of fancy controls and touchscreens). The cheapest thermocycler on eBay goes for $230, but it’s from 1997. These prices are absurd because, again, thermocyclers really haven’t changed much in the last forty years. On paper, then, selling a photonic PCR for $150 should be easy.
But cheap lab hardware has been tried before, and the results are not encouraging. In 2010, a biohacker named Tito Jankowski co-founded a nonprofit called OpenPCR that aimed to build, test, and sell cheap thermocyclers. Each device cost about $250 to build (surprisingly, the aluminum blocks only cost ~$15) and sold for $500. They ended up selling about a thousand machines, and the project is now defunct.
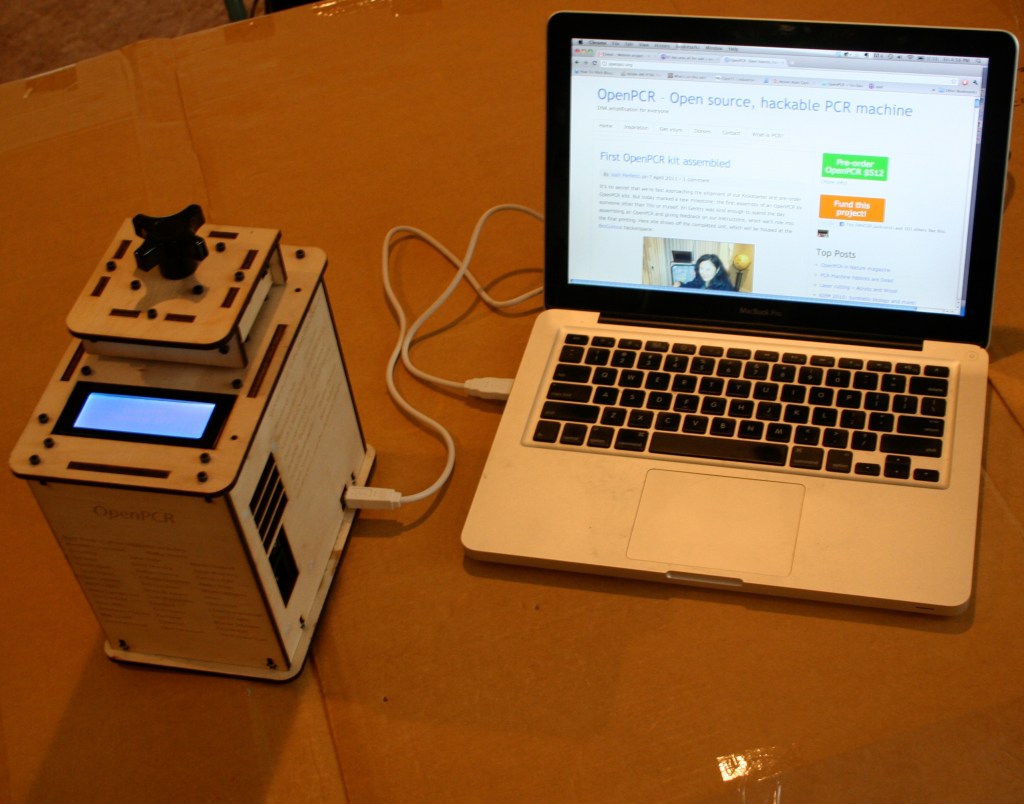

An OpenPCR device, circa 2011.
Before Tito tried selling PCR machines, he made and shipped blue light illuminators — those lights used to visualize gels and “see DNA” after electrophoresis. These blue lights cost about $2,500 from the big companies, but Tito figured he could build one for about $10 and sell it for $100. So he did that, only to find that scientists didn’t want to buy the lights, even though they were nearly identical to the commercial options and cost ~25-times less. Scientists assumed something must be wrong with something so cheap, Tito says. When he doubled the prices, sales went up.
Tito thinks the same idea applies to PCR machines, or really to any lab hardware. Scientists don’t always trust cheap alternatives, even when the data shows they work just as well. There’s also a gap between scientists and engineers; scientists often worry they’ll have to troubleshoot their protocols on an unfamiliar device, or that their experiments will stop working, whereas engineers are often more open to experimenting with, or building their own, hardware.
The problem isn’t only about price, then; there is also a switching cost! The further a new tool strays from what scientists already know, the less likely they are to adopt it. Swapping Taq for Phusion has a low switching cost, because scientists can just replace one enzyme in their PCR for another, without the need to change protocols or use a different thermocycler. The time savings are also large.
Asking labs to throw out their workflow and replace it with lasers and oil droplets, just to save 20 percent on PCR times, is probably not going to work. Some groups might adopt this, like core facilities or robotic labs that could run it and troubleshoot at scale, but the switching cost for most groups is just too high, and the advantages too small.
There is a painful irony to all this. Thermocyclers cost thousands of dollars and haven’t changed in decades, and yet cheaper and slightly faster alternatives will still struggle to spread. Once tools become common, they get entrenched and become difficult to dislodge. Anyone building open-source or low-cost lab hardware should keep this in mind.
Reader comments:
JM: I read your blog on PCR today and I think one thing you overlook is being more bespoke.
As you point out, the length of DNA determines extension time. However, this causes an issue when you want to do a 96 well plate of different length pcrs at the same time (common in our work as we pcr as a first step after ordering and before goldengate to increase goldengate efficiency for short fragments). At the moment, we will split plates into similar lengths (at synthesis order or after receiving them) or try and run things at a single extension time that roughly works for everything. As such, we have been looking at the icon96 as each “well” in the machine can be controlled separately allowing individual optimization of wells despite staying in a single piece of labware. This also has benefits if you want to use more than one set of primers with different annealing temperatures. Finally, they have an interesting innovation around the number of cycles. They have added fluorescent dyes to enable stopping when a certain concentration of DNA is reached regardless of starting material. This enables potentially reduced timing on cycles without sacrificing downstream applications.
In short, they seem to be offering on the go protocol optimization at the single well level.
- My reply: I had no idea that such a technology existed. Thank you for writing and sharing this with me; it sounds very cool! I’ll look into it more closely.
HL: Without systems level engineering, bio processes, hardware, and ecosystems will be resistant to improvement.
PCR itself can absolutely be faster- cut cycles. Those that do qPCR amplification for custom NGS can see near saturation far under 30 cycles. The overlooked question is how much product is enough? We often have far too much. How much did you start with? What’s the ratio of input to primer concentration?
Many seemingly mild improvements across the system will yield productive outcomes, particularly in the coming age of autonomous labs.
SD: Perhaps one effective way to win over skeptical scientists’ hearts is to let them try a lower-cost instrument for free (say, for a year). The lower production costs of these machines may make such an approach very feasible.
MS: It’s not the length of DNA itself, it’s the speed of the polymerase. Biology operates at its own pace. And the optimal rate of the polymerase(s) has been well-characterized. Ramp speed is interesting. You can (and some have) sped that up by doing PCR in capillary tubes (thinner volumetric cross-section) in a hot-air cycler (instant temperature change outside) e.g., Roche’s early LightCycler took 30 minutes.
But the 32 sample capillary system can’t scale with hot air. It was optimizing for the wrong thing. Labs often want to scale for throughput with not just 96 well plates but 384 well plates. Why worry about cutting cyling time in half when you can get 10 times as many samples only taking twice as long. It’s not like there isn’t something else to get done in the lab during a PCR run.
RA: With speeds of engineered polymerases at 5s/Kb, I’m not sure that enzyme yield is the bottleneck. The enzyme has still to disengage and primers to anneal, enzyme find the template again, etc, and reaction gets self-inhibited over time.
Maybe we got to think out of the box. E.g. a rolling circle polymerase that works at temperatures so high it does not need strand displacement, because the template dissociates right after each nt extension. That could be very fast. Or a way to dynamically control the amount of dna product and reaction components, like a microfluidic device which keeps mixing channels, would tackle the issue of saturation and keep the reaction continuously exponential. Maybe a magic polymerase which instead of just copying the template ones, it creates multiple copies in parallel at each cycle (e.g. adds nucleotides simultaneously at different positions, and amplifies with a combination of regular amplificstion and strand displacement). Or maybe a rolling circle on a concatemerised template, which at each cycle it makes more than 2x the initial template… I.e. ideas to brainstorm, throw in the bin and keep the gold.
- My reply: Excellent comment; thanks for the insights. I had no idea a polymerase could move at 5 kb/s, because I’d imagine the diffusion of nucleotides would become rate-limiting, especially over later cycles.
EM: I remember setting up my first PCR in 2014 and having to add all the DNTPs and shit, took forever. Actual nightmare and annoying. [Master Mix] efficiency gains were unreal…. I’m trying to think clinically, I’m not sure I can think of any circumstances where PCR is time sensitive enough where an hour here or there makes a difference. For bacterial differentiation they do MALDI-TOF iirc, for sensitivites and stuff, and PCR is for identification of specific organisms sometimes (stool, urethral swabs etc) but empiric treatment is generally started first for a lot of these.
MPS: Photonic PCR might be quite good for qPCR since that’s often read out using fluorescence, so having optics integrated would help. Or ddPCR.
PC: Interesting article. There are now much faster polymerases out there that could cut down on the proportion of the reaction committed to extension time (we use Repliqa – as fast as 1s/kb), making ramp rate savings proportionally higher.
FC: It’s worth looking at what companies like MBS/NextGenPCR have been doing with SBS format plates (e.g., 3 temp 40 cycle PCR in 2 mins) – many things can be done to make PCR go quite a bit faster, especially qPCR (if you need access to post-reaction material your options change quite a bit), and even for long amplicons – and it’s entirely possible there’s a boatload of opportunity in enzyme engineering (the Roche folks are incredible at this)